Introduction
The Anti-Amyloid Treatment in Asymptomatic Alzheimer’s Disease (A4) study (ClinicalTrials.gov ID: NCT02008357) was a phase 3 randomized, double-blind, placebo-controlled clinical trial designed to evaluate the efficacy of solanezumab in slowing cognitive decline among cognitively unimpaired individuals with elevated brain amyloid levels.1 From over 6,700 individuals screened, 1,169 participants aged 65 to 85 were enrolled and randomized. The trial, conducted between 2014 and 2023, included a 240-week blinded treatment period followed by an open-label extension. Participants underwent comprehensive assessments every six months, including clinical evaluations, cognitive testing, neuroimaging, and biospecimen collection. A companion observational study, the Longitudinal Evaluation of Amyloid Risk and Neurodegeneration (LEARN) (ClinicalTrials.gov ID: NCT02488720), enrolled 538 individuals who did not meet A4 eligibility criteria due to low amyloid burden and followed a similar longitudinal assessment protocol.
The A4 and LEARN studies adhered to NIH data-sharing policies and further embraced the principles outlined by the Collaboration for Alzheimer’s Prevention.2 Accordingly, data were released in two phases to promote transparency and accelerate scientific discovery while maintaining strict safeguards for participant privacy. The initial release, comprising screening and baseline neuroimaging data, occurred in 2018 and 2020. The complete longitudinal dataset was made available in 2024.
In a previous publication, we presented the overarching strategy for data sharing adopted by the A4 and LEARN studies.3 In the present article, we build upon that foundation by providing a detailed account of the methodologies, platforms, and outcomes associated with disseminating neuroimaging data collected throughout the studies. We describe the phased release of de-identified datasets, the infrastructure supporting data access, and the collaborative framework that guided these efforts. Furthermore, we assess the impact of these data-sharing initiatives on secondary research and highlight key lessons learned to inform future large-scale neuroimaging studies.
Methods
Imaging studies
Magnetic resonance imaging
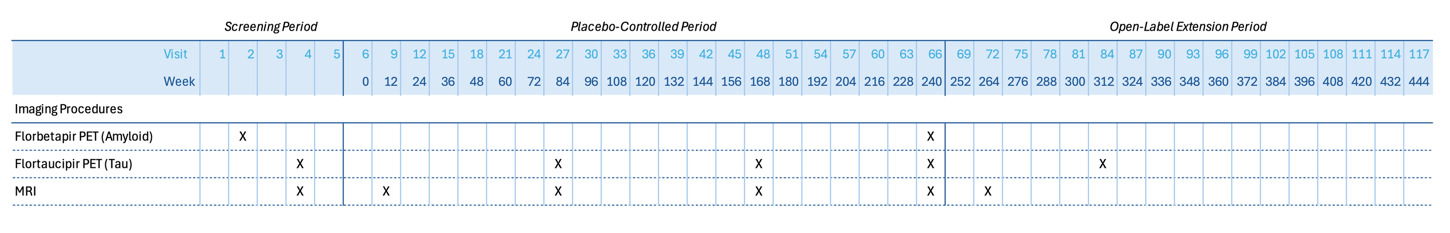
Magnetic resonance imaging (MRI) was used in the A4 and LEARN studies to obtain high-resolution images of brain structure and function. MRIs were collected at scheduled intervals and when clinically indicated, following standardized procedures implemented by qualified imaging sites using the A4 MRI protocol. This protocol was centrally developed from the ADNI MRI guidelines and electronically distributed to sites prior to scanner qualification. All scans were obtained on 3 Tesla (3T) systems equipped with multi-channel head coils (additional details may be found in the Supplemental Materials). Each scheduled session included six sequences: T1-weighted MPRAGE, FLAIR, T2 Spin Echo (SE), T2, diffusion-weighted imaging (DWI), and resting-state functional MRI (fMRI). Structural sequences (T1 MPRAGE, FLAIR, T2 SE, T2, and DWI) were used during screening to exclude participants with significant vascular pathology, microhemorrhages, or other incidental findings, and post-enrollment to monitor for amyloid-related imaging abnormalities (ARIA) and derive secondary outcomes such as brain volumetrics and white matter hyperintensities. Resting-state fMRI was used to generate functional connectivity maps as a secondary outcome measure. Follow-up MRIs were conducted at weeks 12, 84, and 168, or at early termination, to assess the emergence of ARIA-E (edema) and ARIA-H (hemorrhage), with asymptomatic cases permitted to continue treatment (Figure 1). Volumetric analyses were performed to evaluate changes in brain volume, and functional connectivity analyses assessed network integrity over time. Participants who completed the blinded phase were invited to join an optional open-label extension, during which additional MRIs were collected at weeks 264 and, in some cases, 312 (Figure 1).
Amyloid positron emission tomography with florbetapir
In the A4/LEARN studies, amyloid positron emission tomography (PET) imaging of the brain was performed using the A4 PET scanning protocol. Baseline PET scans were used to determine eligibility based on amyloid burden. Follow-up PET scans were collected at 240 weeks or early study termination (Figure 1). Before each scan, participants received a single intravenous administration of approximately 370 MBq (10 mCi) of florbetapir F 18 (FBP), a tracer known to bind to fibrillar amyloid plaques, about 50 minutes before sequence acquisition. The PET acquisition procedure typically lasted 20 minutes with a target window of 50-70 minutes after injection with the radiotracer (additional details may be found in the Supplemental Materials). Changes in brain amyloid burden, assessed by florbetapir binding and measured by mean cortical summary uptake value ratio (SUVR), were compared between solanezumab- and placebo-treated participants who underwent baseline and endpoint (240 weeks) florbetapir scans.
Tau positron emission tomography substudy with flortaucipir
In a subset of A4/LEARN participants, the PET tracer 18F-AV-1451 (also known as 18F-T807 and branded as flortaucipir (FTP)), noted for its high affinity to paired helical filament tau deposits, was utilized to investigate whether anti-amyloid therapy could slow the progression of tau pathology over time. 390 A4 participants with preclinical Alzheimer’s disease (AD) underwent sequential PET scans at three time points to monitor changes in tau deposition: baseline, 84 weeks, and 240 weeks or at early study termination (Figure 1). Before each scan, participants received a single intravenous administration of approximately 370 MBq (10 mCi) of FTP administered as a single IV bolus followed by a 15-20 mL saline flush, with imaging initiated approximately 75 minutes post-injection. The PET acquisition procedure typically lasted 30 minutes, targeting a standardized 75-105-minute post-injection window (additional details may be found in the Supplemental Materials). The study also examined the association between increasing tau levels and cognitive decline in the placebo group, aiming to identify a potential biomarker of disease progression. This imaging component was included as a protocol addendum, as not all study sites had access to the 18F-AV-1451 tracer. A similar protocol was implemented in the LEARN study (n = 55), with tau PET scans collected at baseline, 72 weeks, 168 weeks, 240 weeks, or at early termination.
Data acquisition
Ethical approval was obtained from the institutional review board or ethics committee at each participating site. All participants and their study partners provided written informed consent prior to study initiation, including consent for data collection and data sharing.
Site qualification
The A4/LEARN studies were conducted at 68 trial sites in Australia, Japan, and North America. As part of the site start-up process, each site’s MRI and PET scanners underwent an initial qualification process that included the installation of study-specific protocols and settings. This process also included the acquisition of a qualification scan using an imaging phantom. Each qualification scan was uploaded to the A4/LEARN data management system4 and assessed for compliance with study imaging protocols by the modality-specific imaging laboratory. Scanners that passed quality control were deemed certified for use in the trial. Occasional re-certification processes were initiated by the imaging laboratory or trial sites based on 1) updates to the scanner software or settings, 2) imaging protocol updates, or 3) the introduction of a new scanner.
Image collection
Participant scans were acquired according to the designated study- and modality-specific protocols. After acquisition, site personnel relabeled each scan with the appropriate participant ID and uploaded it to the A4/LEARN data management system. Imaging data were securely stored, backed up, and archived according to standard procedures. To ensure consistency, sites were encouraged to submit scans in the Digital Imaging and Communications in Medicine (DICOM) format. Although most sites followed this guidance, a small number of scans were submitted using an older format known as Emission Computerized Axial Tomography (ECAT), which required conversion to DICOM to maintain standardization across the dataset.
Once uploaded, each scan was registered in the imaging database, where it underwent an initial series of quality control (QC) checks to verify data integrity and image quality. To safeguard participant privacy, a predefined set of DICOM header fields was de-identified. This process removed all personally identifiable information (PII) while retaining essential metadata required for downstream QC and analysis.
Image quality control, processing, and analysis
The A4/LEARN study implemented a comprehensive QC framework for MRI and PET imaging to ensure data integrity, reproducibility, and compliance with regulatory standards. These procedures were developed in collaboration with leading academic and industry experts and were critical for harmonizing imaging data across multiple sites and scanner platforms. The QC strategy addressed both pre-acquisition and post-acquisition phases, incorporating site qualification, standardized acquisition protocols, automated QC checks, and centralized review processes.
MRI scans that did not meet quality control standards and were required for safety review were re-acquired to ensure participant safety and protocol compliance, while scans not needed for safety review such as resting-state fMRI were generally not repeated if they failed QC. In contrast, PET scans were rarely re-acquired due to radiation exposure concerns, except in exceptional cases where re-acquisition was deemed essential and approved by study leadership.
The study team collaborated closely with several academic imaging experts to facilitate the sharing of analytic datasets derived from the MRI and PET data. These included MRI Safety Reads to identify microhemorrhages and superficial siderosis, MRI volumetric measures, amyloid PET eligibility measures (SUVr, clinical read), standardized amyloid and tau PET SUVr measures.
MRI quality control, processing, and analysis
Before participant enrollment, each imaging site underwent rigorous MRI Site Qualification to confirm scanner performance and adherence to protocol requirements. This process typically involved scanning an ADNI phantom using A4-specific sequences provided by the Mayo Clinic Aging and Dementia Research (ADIR) laboratory, including Localizer and QC Phantom Sagittal 3D Accelerated MPRAGE/IRSPGR.5 Sites with scanners not previously used in multi-center trials or with poor phantom results were required to perform human volunteer scans. The ADIR QC team reviewed all submissions to verify parameter accuracy, image quality, and scanner performance. Only one scanner per site was qualified for A4, and all subsequent scans had to use that scanner to maintain longitudinal consistency. Sites failing QC were instructed to re-scan after corrective actions.
Following acquisition, 3D T1-weighted MRI scans were uploaded to the imaging database and processed using NeuroQuant (NQ), an FDA-cleared automated segmentation tool.6 The pipeline began with automated QC to detect motion artifacts and other anomalies, followed by generation of a personalized atlas for each participant, adjusted for age and sex, to guide segmentation. NeuroQuant then performed automated segmentation and computed regional brain volumes, including the Hippocampal Occupancy (HOC) Score - a sensitive biomarker of medial temporal lobe atrophy relevant to AD progression. The HOC Score was defined as the ratio of hippocampal volume to the combined volume of the hippocampus and inferior lateral ventricles, serving as a robust proxy for neurodegenerative changes. All volumetric measures were archived for downstream analyses and shared with collaborating investigators to support secondary research.
PET quality control, processing, and analysis
All PET imaging operations were managed by Molecular NeuroImaging LLC (MNI), later acquired by Invicro. Responsibilities included qualifying imaging centers, providing technical support, and overseeing data quality control and analysis. Prior to participant recruitment, each imaging center underwent a technical setup that assessed scanner hardware and software, quality assurance procedures, and acquisition protocols using a Hoffman 3D phantom. Standardized documentation was provided through a Technical Binder. FBP and FTP PET data were reviewed for compliance, analyzed using validated algorithms, and archived in their original format. Data processing and quantitative analysis were performed on π.PMOD workstations, with backups maintained according to standard operating procedures. All imaging data were retained for fifteen years unless otherwise specified.
PET image processing and quantitative analysis were carried out by an Image Processing Specialist (IPS) using π.PMOD software validated for 21 CFR Part 11 compliance, including full audit trails. Analyses were blinded to treatment status and followed a standardized pipeline that included motion correction across the X, Y, and Z planes, frame averaging, and normalization to Avid Template Space (Avid Radiopharmaceuticals Inc.).7,8 Normalization involved a two-step process: an initial rigid alignment to the Avid PET reference template, followed by brain normalization using predefined parameters with a 3-mm sampling rate and a linear method. All transformations were verified during quality control, and each processing stage generated traceable files to ensure data integrity and reproducibility. Only scans that passed quality control were included in downstream analyses.
For Florbetapir F 18 PET imaging, standardized uptake values (SUVs) were calculated using the formula where is the tracer concentration, is the injected dose, and is the participant’s body mass. Participant-specific data, including weight and dose, were recorded at the time of injection and entered into the imaging database using double data entry to ensure accuracy. These data were imported into π.PMOD software for SUV computation on spatially normalized PET images. A trained and annually certified IPS applied a predefined Volume of Interest (VOI) template, referred to as the Avid Eligibility VOI Template (Avid Radiopharmaceuticals Inc.),7,8 which encompassed seven brain regions. VOI placements were quality-checked for anatomical accuracy, particularly in cases of brain atrophy. Once verified, the VOI template was applied to extract SUV values, which were saved as non-editable text files. π.PMOD was then used to compute standardized uptake value ratios (SUVRs) for the following brain regions: medial orbital frontal (XLAAL_FRONTAL_MED_ORB), parietal (LNEW_PARIETAL), temporal (NEW_TEMPORAL_2), precuneus (LPRECUNEUS_GM), posterior cingulate (LLPOSTERIOR_CINGULATE_2), and anterior cingulate (LANTERIOR_CINGULATE_2), using the mean uptake in the whole cerebellum (BLCERE_ALL) as the reference region.8,9 A composite SUVR was also calculated as the unweighted average of the six cortical VOIs listed above, excluding the reference region.10 In cases where it was not possible to obtain SUVs from the FBP PET images, SUVRs were calculated using the units in which the data was submitted. All SUVRs underwent review by a nuclear medicine physician prior to submission to the study database.
SUVRs for FTP were derived using an MNI template space atlas of the GTMseg atlas from11 that was created using data from the baseline cohort of the Harvard Aging Brain Study (HABS). Briefly, for each of 280 baseline T1 MPRAGE images, the native space GTMseg atlas produce by PETsurfer was mapped to MNI space using a diffeomorphic transform computed with the SPM12 unified segmentation/normalization algorithm. Each voxel in MNI space was then assigned the GTMseg label that most frequently occurred across all 280 baseline HABS scans to generate a template space GTMseg atlas. The A4 FTP data were also normalized to MNI space by coregistering the FTP scans to the corresponding T1 image and then applying an MNI space transform that was derived from the T1 image. Once the FTP-PET images were transformed to MNI space regional measurements were obtained for all 100 regions defined in the standard GTMseg atlas, and SUVRs were generated by normalizing the regional measurements to the PET signal in the cerebellar grey reference tissue.
Preparing data for sharing
The A4/LEARN investigators established a systematic data preparation pipeline to ensure the imaging data’s consistency, privacy, and usability. This process included reconciling external QC findings, removing personally identifying information (PII/PHI), converting raw DICOM files to the more portable Neuroimaging Informatics Technology Initiative (NIfTI) format using dcm2niix,12 and organizing the JSON header metadata according to the Brain Imaging Data Structure (BIDS) standard13 to enhance interoperability and accessibility.
The NIfTI-formatted imaging studies were subsequently de-faced/re-faced using the mri_reface tool.14,15 This tool, which has been demonstrated to provide strong participant privacy protections while retaining the scientific utility of the data, was used to de-face/re-face both MRI and PET sequences. For MRI, a subset of the sequences were de-faced/re-faced: 1) T1 - Sagittal 3D Accelerated MPRAGE, 2) T2_star - Axial T2-Star, 3) FLAIR - Axial T2-FLAIR, and 4) T2_SE - Axial T2-TSE with Fat Sat. To ensure the quality of the final de-faced/re-faced images, a 100% expert quality review was conducted.
Data sharing platforms
Baseline data were shared through the Imaging & Data Archive (IDA) (ida.loni.usc.edu), which is managed by the Laboratory for Neuro Imaging (LONI) at the Mark and Mary Stevens Neuroimaging and Informatics Institute at the University of Southern California (USC).
To broaden access to the final longitudinal dataset and meet investigator communities where they are, the A4/LEARN investigators chose to distribute the data across multiple data platforms. These included LONI IDA, which primarily attracts researchers with a focus on neuroimaging; Sage Bionetworks’ Synapse platform (www.synapse.org), which is widely used by the omics research community; and a custom-built platform developed in collaboration with Gates Ventures (www.a4studydata.org), designed to support general data usability and broader data exploration and integration.
Measuring scientific impact
To measure the scientific impact of the A4/LEARN data-sharing efforts, investigators conducted a bibliometric analysis using Google Scholar (scholar.google.com), Dimensions AI (app.dimensions.ai), and Web of Science (webofscience.com). In accordance with the A4 Data Use Agreement (DUA), which requires researchers to acknowledge the A4/LEARN study in their publications, Google Scholar and Dimensions AI were queried using the phrase ‘a4 study atri public-private-philanthropic’. Web of Science was searched using the terms ‘a4 study’, ‘a4 trial’, ‘a4 clinical trial’, and ‘a4 clinical study’ across all fields, with ‘alzheimer’ specified as a topic keyword. Results from all platforms were merged, restricted to peer-reviewed articles and registered preprints, and filtered to include publications dated between 2020 and 2025 (inclusive). Each publication’s author list was reviewed to determine attribution to the A4/LEARN study team, and each abstract was analyzed to assess the study’s objectives, design and methods, key findings, and conclusions. A thematic analysis was then conducted to identify overarching themes and research trends within the selected literature.
Results
Image collection characteristics
Baseline dataset
The key characteristics of the A4/LEARN baseline image collection are summarized in Table 1. This collection includes screening and pre-randomization MRI (n=1771) and amyloid PET (n=4468) scans used to assess study eligibility and baseline tau PET sequences (n=449). A large number of images collected from participants who failed to meet the screening criteria for both studies are also included.
Final study dataset
The key characteristics of the final A4/LEARN study image collection are summarized in Table 2. The dataset includes 5,661 amyloid PET scans and 1,576 tau PET scans, making it one of the largest high-resolution longitudinal preclinical AD PET imaging collections currently available for secondary research. The distribution of longitudinal MRI and PET images by study phase is shown in Table 3, and participant counts by number of longitudinal scans, study, and scan type are detailed in Table 4. Longitudinal MRI, amyloid PET, and tau PET images were further quantified by days since each participant’s initial consent date in the A4 study (Figure 2) and the LEARN study (Figure 3). For MRI, the most common scanner manufacturers were Siemens (67.0%; models: Skyra, Trio, Verio, Biograph), GE Medical Systems (23.5%; models: Discovery, Signa), and Philips (8.6%; models: Achieva, Ingenia, Intera). For amyloid PET, Siemens accounted for 50.9% (Biograph, SOMATOM), GE Medical Systems for 21.0% (Discovery), and Philips for 17.1% (Ingenuity, Gemini, Guardian, Vereos). For tau PET, Siemens represented 42.6% (Biograph, SOMATOM), Philips 20.5% (Gemini, Ingenuity, Guardian, Vereos), and GE Medical Systems 18.2% (Discovery).
Contributed analysis datasets
After the initial sharing of the imaging data, several academic teams have collaborated with the A4/LEARN investigators to provide additional analytic datasets generated using novel methods. These include adjusted tau PET SUVR measures using PetSurfer partial volume correction, the Stanford PET pipeline,16 and retinal Amyloid measures.17
Data use requests
Baseline imaging data
Baseline tabular and imaging data from the A4 and LEARN studies were released in December 2018 and early 2020, respectively. The tabular dataset included clinical, biomarker, neuropsychological, laboratory, and imaging measures, while the imaging data comprised re-faced MRI, amyloid PET, and tau PET scans. As of May 2025, investigators from more than 60 countries have submitted 1,907 data access requests, of which 1,741 (91.3%) were approved (Figure 4). Since then, 179 approved users have collectively downloaded over 1.2 million MRI and PET imaging sequences (Figure 5).
Final study imaging data
Final longitudinal tabular and imaging data from the A4 and LEARN studies were released in July 2024. As of May 2025, investigators from 30 countries have submitted 397 data access requests, of which 379 (95.5%) were approved (Figure 6). Among these approved requests, 347 (91.6%) were processed through the Gates Ventures platform and 32 (8.4%) through the Synapse platform.
Scientific impact
A bibliometric search conducted on May 30, 2025, yielded the following results: 80 records from Google Scholar, 91 from Web of Science, and 91 from Dimensions AI. Of these, 19 publications were attributed to the A4/LEARN study team. These results include several highly cited imaging-related articles with more than 70 citations that have advanced the understanding of the natural history of preclinical AD, the topographic patterns of tau PET, and changes in plasma markers of AD, as well as the demographic nuances of imaging markers.16,18–20 These publications have also informed revisions to diagnostic and characterization guidelines for AD based on biomarker criteria, established the foundations of recruitment science in preclinical AD trials, and improved understanding of the impact of amyloid PET result disclosure at scale.21–23 Notably, the majority of these articles referenced the baseline dataset.
More broadly, the A4 and LEARN data have made substantial contributions to advances in preclinical AD research, underscoring the increasing precision in identifying risk, tracking pathology, and optimizing trial design. These studies have demonstrated how genetic factors such as APOE4 homozygosity and protective APOE2 and KL-VS variants influence disease trajectories, with APOE4 homozygotes exhibiting near-complete penetrance of AD biomarkers and earlier symptom onset, which has important implications for therapeutic development.16,18,19,24–31 They have also shown how blood-based biomarkers, including plasma P-tau217 and Aβ42/Aβ40 ratios, together with predictive algorithms and tools such as PARS and AmyloidPETNet, are improving early detection and reducing reliance on costly imaging.29,32–37 Furthermore, these studies have highlighted that cognitive and functional changes, including subtle impairments in memory and daily activities, can be detected through tools such as Cogstate Brief Battery (CBB), Computerized Cognitive Composite (C3), and Stages of Objective Memory Impairment (SOMI) staging, while subjective and partner-reported assessments provide complementary insights.19,38–43
In addition, research based on these data has clarified how lifestyle factors such as sleep, physical activity, and moderate alcohol consumption affect cognition through distinct mechanisms, and how psychological responses to biomarker disclosure underscore the need for ethical communication.44–53 Studies have also emphasized the importance of inclusive research by revealing diverse genetic architectures and disparities in biomarker expression across racial and ethnic groups.20,24–28,31,42,54,55 Finally, recent work has explored innovations in imaging and scalable screening methods, including retinal fundus imaging and harmonized cognitive composites, which are enhancing trial efficiency and broadening access to early intervention strategies.17,34,43,56–58
Discussion
As discussed in,3 our results provide further evidence of the feasibility and scientific utility of broad and timely sharing of AD trial data, even for trials such as A4 that failed to show a therapeutic benefit.1 The growing number of high-impact articles derived from these shared data provide additional supporting evidence, and are critical for informing future trials. To date, most analyses have focused on the cross-sectional baseline dataset. With the recent release of the final longitudinal dataset, we anticipate a wave of new research findings that will enhance our understanding of disease progression and therapeutic mechanisms.
While interest in the baseline data remains strong, with a steady stream of new data use requests, demand for the final longitudinal dataset is growing faster. For instance, the final dataset reached 250 approved requests in just six months - four months sooner than it took the baseline data to achieve the same milestone, which required a full year. This trend highlights the potential utility and relevance of the final dataset for the AD research community.
We are encouraged to see our efforts to promote open science and data sharing in Alzheimer’s clinical trials via our A4/LEARN experience have vitalized other investigators to adopt similar principles and share the results of their analyses, employing the A4 repository as a trusted hub to broadly disseminate newly derived neuroimaging data and measures generated using novel methods to the AD research community. We plan to continue our work to develop data-sharing approaches and platforms to facilitate these activities, establishing processes to ensure high-quality data submissions and acknowledging contributors for their efforts to enrich the repository.
Limitations
While our efforts to share the A4/LEARN data have been successful, several limitations remain. First among them is the challenge of accurately quantifying the impact of our data-sharing activities. Our current method - relying on multiple literature search tools - offers only a rough approximation. We are exploring more robust strategies to address this issue, such as assigning Digital Object Identifiers (DOIs) to each dataset or data release. This approach would enhance citability, facilitate more precise data usage tracking, and ultimately provide a clearer picture of the scientific contributions enabled by these shared resources.
Second, key barriers remain for investigators seeking to integrate clinical and neuroimaging data with non-imaging data types such as genome-wide association studies (GWAS) or whole-genome sequencing (WGS) data. Due to the heightened privacy risks associated with genetic information, these datasets are stored in separate, highly controlled repositories that are not yet interoperable with our current data-sharing platforms. Addressing these integration challenges is a priority for future development efforts.
Finally, the substantial scientific interest and utility of the A4/LEARN data justified the significant resources required to support its sharing. To ensure sustainability and equity, both immediate and long-term costs associated with data sharing should be explicitly incorporated into clinical trial budgets. Dedicated funding for infrastructure, maintenance, and ongoing accessibility is essential to scale these efforts while preserving quality and usability over time. Encouragingly, the 2023 NIH Policy for Data Management and Sharing (NOT-OD-21-013), effective January 25, 2023,59 now requires data management and sharing plans, along with associated budgets, to be included as part of grant applications.
We maintain that open access to clinical trial data is essential for promoting transparency, reproducibility, and trust in the scientific process. It also maximizes the scientific return on investment by enabling broader secondary use. However, in response to these questions, we continue to refine our processes to improve efficiency and sustainability.
For example, we have identified opportunities to automate tasks that were previously manual, such as reducing the need for manual quality control during the de-facing process. These improvements are expected to yield meaningful cost savings and enhance scalability. Additionally, we are exploring the use of “crowdsourcing” strategies to help distribute the long-term costs of maintaining data repositories, which may offer a viable path toward sustainability. We plan to report on these efforts in future work.
Conclusions
Clinical trials rely on the contributions of a diverse group of stakeholders, including study participants and their partners, investigators, research sites, sponsors, regulators, and the broader public. A successful trial - regardless of whether its outcome is positive or negative - provides a definitive answer to the primary research question it was designed to address. By sharing trial data widely, we can foster trust in the scientific process through increased transparency and reproducibility. Moreover, broad data sharing extends the value of these investments by enabling the investigation of a wider range of research questions than originally envisioned by the study team, thereby enhancing the overall scientific utility of the data.
In the case of the A4/LEARN neuroimaging datasets, we anticipate that these data will not only advance research in AD but also contribute to broader studies on aging and other neurological conditions, further increasing their long-term impact.
Data and Code Availability
The A4/LEARN data package, which includes the complete final longitudinal datasets from the A4 and LEARN studies, is available to qualified researchers through a controlled-access process at https://www.a4studydata.org/, subject to institutional approvals, data use agreements, and applicable regulatory and ethical requirements. Owing to participant privacy considerations, the data are not publicly available. The data package also includes an R package that enables replication of the primary study results. In addition, investigators may access the alzverse R package (https://atri-biostats.github.io/alzverse/; Donohue et al., 2026; https://doi.org/10.1002/alz.71152), which facilitates interaction with the A4/LEARN datasets.
Funding sources
The A4 and LEARN Studies were supported by a public-private-philanthropic partnership which included funding from the National Institute of Aging of the National Institutes of Health (R01 AG063689, U19AG010483 and U24AG057437), Eli Lilly (also the supplier of active medication and placebo), the Alzheimer’s Association, the Accelerating Medicines Partnership through the Foundation for the National Institutes of Health, the GHR Foundation, the Davis Alzheimer Prevention Program, the Yugilbar Foundation, Gates Ventures, an anonymous foundation, and additional private donors to Brigham and Women’s Hospital, with in-kind support from Avid Radiopharmaceuticals, Cogstate, Albert Einstein College of Medicine and the Foundation for Neurologic Diseases.
Ethical statement
Ethical approval was obtained from the institutional review board or ethics committee at each participating site. All participants and their study partners provided written informed consent prior to study initiation, including consent for data collection and data sharing.
Acknowledgements
The authors would like to thank the entire A4 and LEARN Study Teams, including all site principal investigators, and their dedicated site staff. Special gratitude to the A4 and LEARN participants and their study partners, without whom these studies would not be possible.
Conflicts of interest
GJM has received research support from the National Institutes of Health (NIH), the Alzheimer’s Association, the American Heart Association, Gates Ventures, Eli Lilly, and Eisai. APS: Nothing to report. MD: Nothing to report. HQ: Nothing to report. RAS reports grant support from the National Institutes on Aging, National Institutes of Health, Alzheimer’s Association, GHR Foundation, and Gates Ventures. She has received trial research funding from Eisai and Eli Lilly for public-private partnership trials. She reported serving as a consultant for AbbVie, AC Immune, Alector, Biohaven, Bristol-Myers-Squibb, Ionis, Janssen, Genentech, Merck, Prothena, Roche, and Vaxxinity. PSA has received grants or contracts from the National Institutes of Health (NIH), Alzheimer’s Association, Foundation for NIH (FNIH), Lilly, Janssen and Eisai and consulting fees from Merck, Biogen, AbbVie, Roche, and Immunobrain Checkpoint.


._ta.png)
.png)
__december_2018_--_may_2025._a_tot.png)
__december_2020_--_ma.png)
_as_of_may_5__2025._cumulat.png)
._ta.png)
.png)
__december_2018_--_may_2025._a_tot.png)
__december_2020_--_ma.png)
_as_of_may_5__2025._cumulat.png)